Biography
Currently, Lingyu Li is a Post-doctoral Fellow at the University of Hong Kong, where she collaborates with Prof. Yuanhua Huang on Bioinformatics. She received her Ph.D. at Shandong University, supervised by Prof. Zhi-Ping Liu. Additionally, She was a joint training Ph.D. student at the University of Hong Kong, supervised by Prof. Wai-Ki Ching.
- Bioinformatics
- Machine Learning
- Biomarker Discovery
- Spatial Transcriptomics
- Single-cell Data Science
- Sparse Statistical Learning
-
PhD in Biomedical Engineering, 2019-2023
Shandong University
-
MSc in Computational Mathematics, 2016-2019
Shandong Normal University
-
BSc in Aplied Mathematics, 2012-2016
Shandong Normal University
Skills
90%
80%
100%
Experiences
Research project:
- Multimodal AI decoding tumor-immune interaction by fusing spatial RNA-seq and histological images
Doctoral thesis title:
- Biomarker discovery methods based on connected network regularized feature selection
Research project:
- Bioinformatics and optimization with applications in biomarker discovery and feature selection
Master thesis:
- Numerical methods and theoretical analysis of a class of groundwater pollution problems
Bachelor thesis:
- Uniform convergence of function term series and its applications
Publications
Paper Overview

In this work, we present CNet_Cox, an connected network-regularized Cox proportional hazards model for identify prognostic biomarkers.
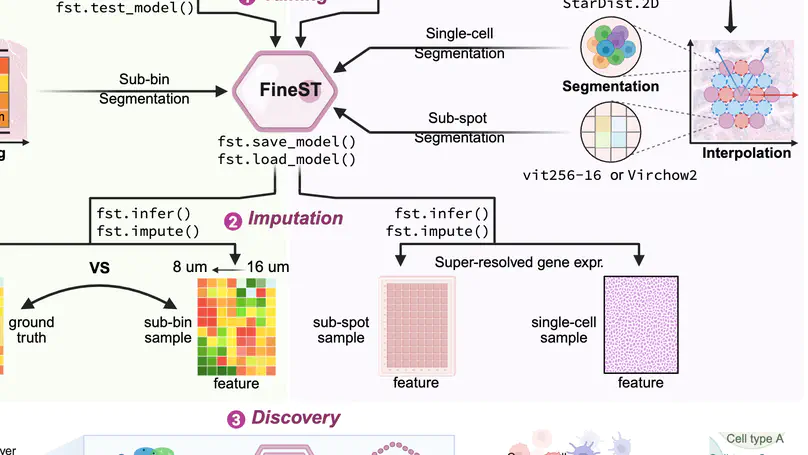
In summary, we have presented FineST, a bimodal contrastive learning framework for fine-grained ligand-receptor identification, which integrates histology images and spatial RNA-seq data to enhance signal strength and achieve synchronized higher resolution.
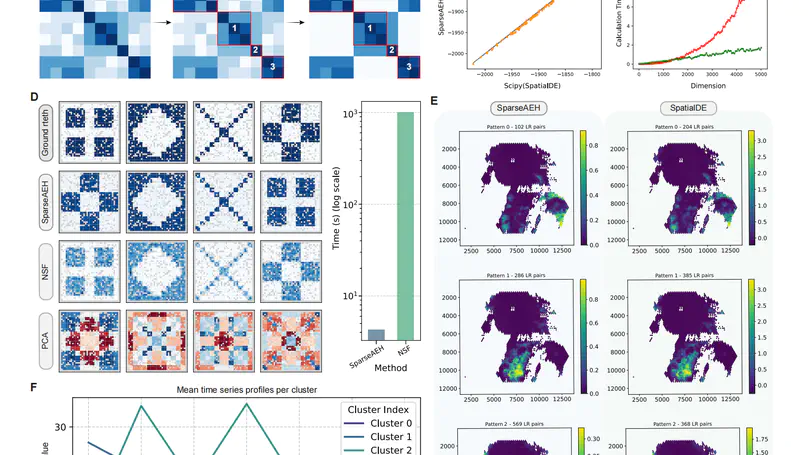
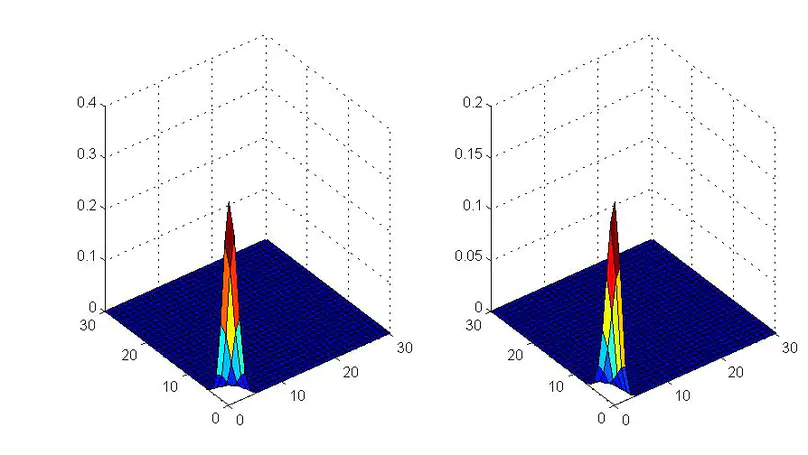
In this work, we present SparseAEH, an EM-algorithm-based method for spatial pattern identification.
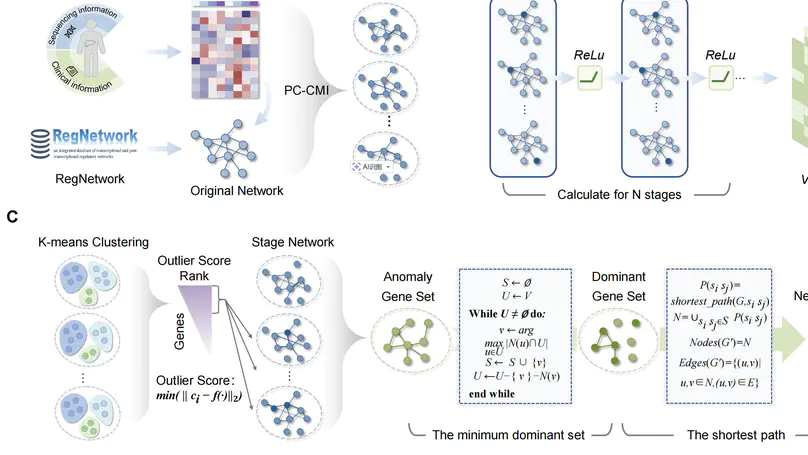
In this work, we offer a mechanism-informed approach to dynamic biomarker discovery, enabling sensitive detection of early-warning signals in HCC with potential translational value.
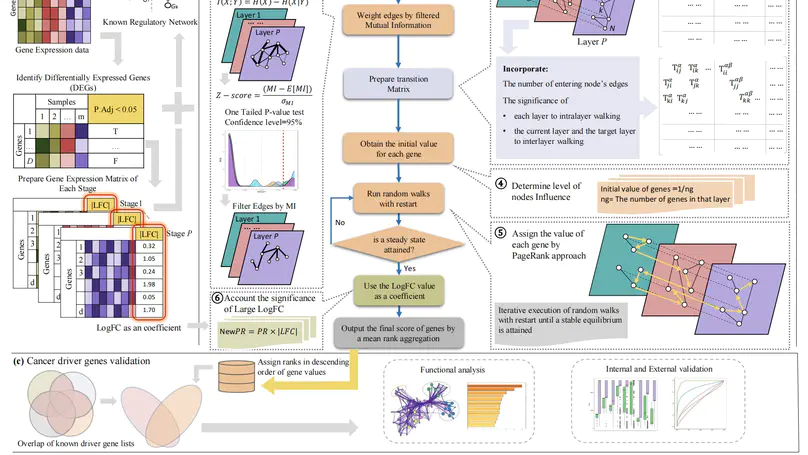
In this work, we present a network-based framework called NetWalkRank, for prioritizing CDGs in the multiplex GRNs with gene expression profiling data.
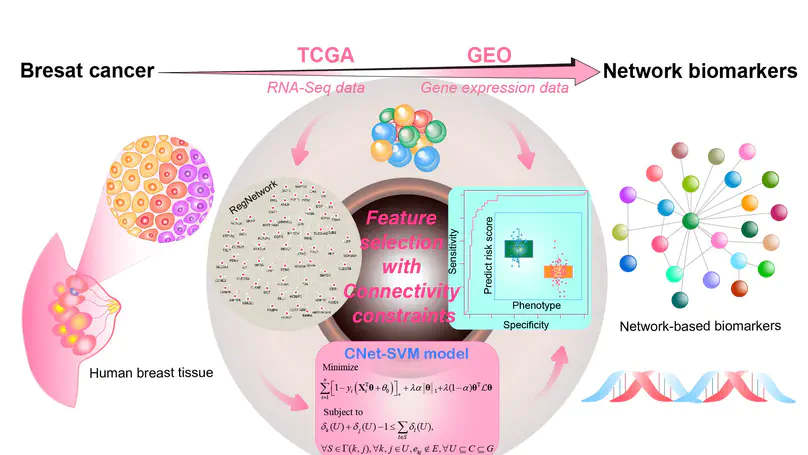
In this work, we provide a connected network-constrained support vector machine (CNet-SVM) model for feature selection considering the structural connectivity in a network.
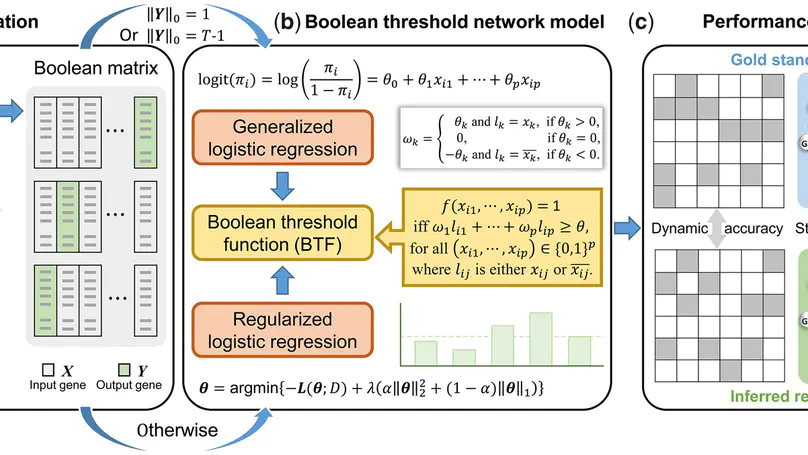
In this work, we provide a computational method of Boolean threshold network (LogBTF) method for gene regulatory network inference from single-cell expression data.
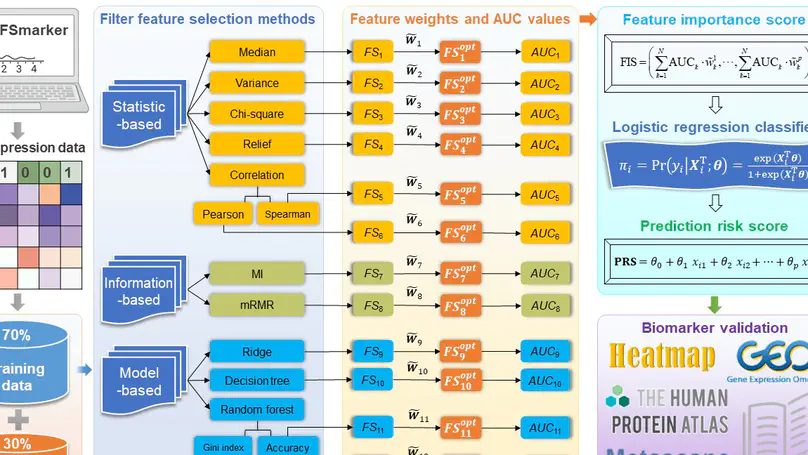
In this work, we provide propose an ensemble feature selection method for biomarker discovery (EFSmarker) based on multiple different independent feature elections methods to give a better approximation to the optimal subset of features.
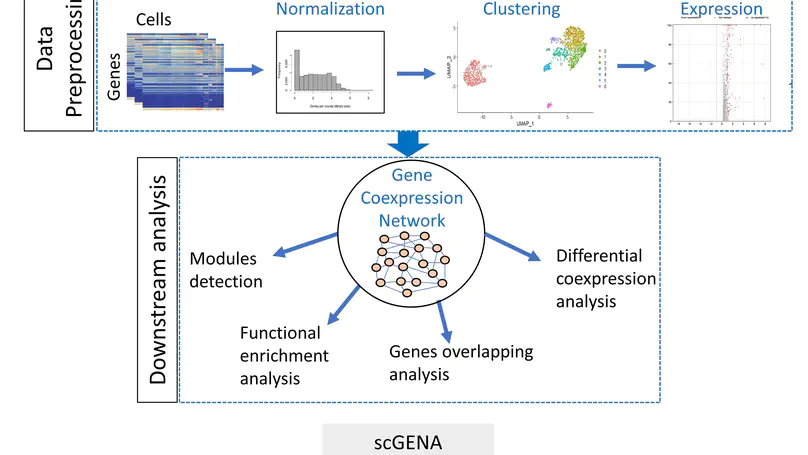
In this work, we provide a single-cell gene coexpression network analysis framework for clustering cell types and revealing biological mechanisms (scGENA).
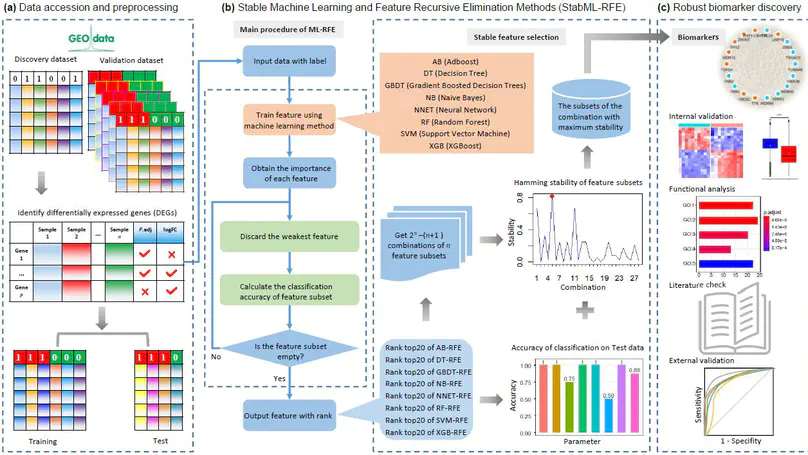
In this work, we provide a computational method of identifying robust biomarkers or signatures from gene expression profiling data by stable machine learning-recursive feature elimination methods (StabML-RFE).
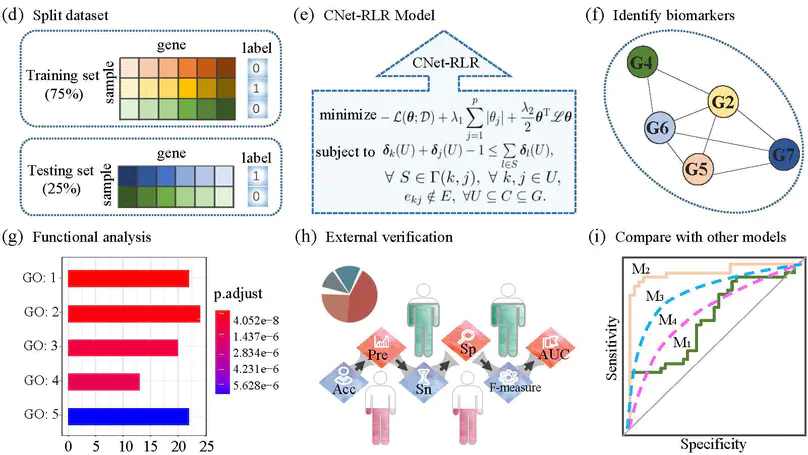
In this work, we provide a computational method of connected network-regularized logistic regression (CNet-RLR) for discovering biomarkers of uterine corpus endometrial carcinoma (UCEC) from genomics data.
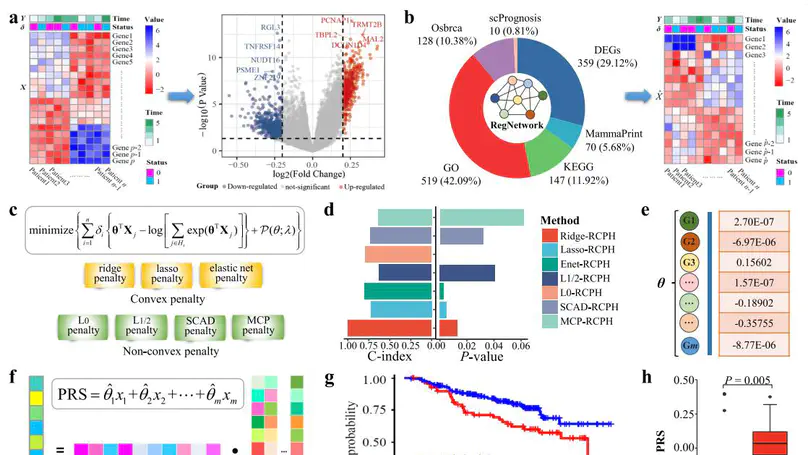
In this work, we provide a computational method of regularized Cox proportional hazards models (CoxReg) for detecting prognostic biomarkers of breast cancer (BRCA) from genomics data.
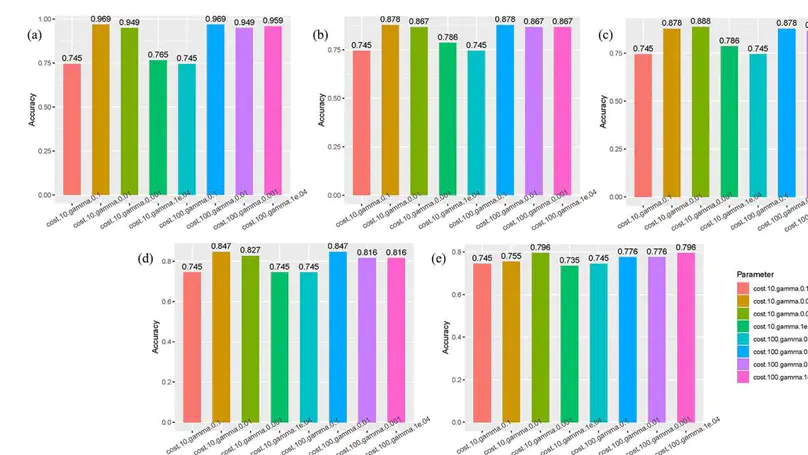
In recent years, the identification of descriptive genes of biomarkers based on gene expression microarray data has attracted much attention in the field of bioinformatics.
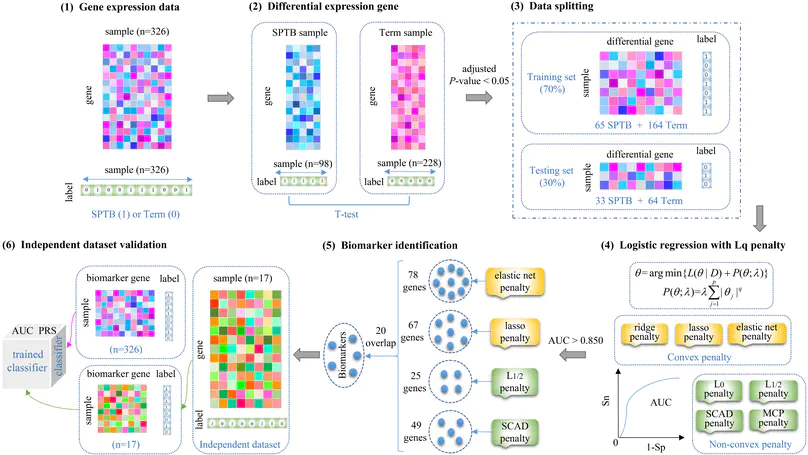
In this work, we provide a computational method of regularized logistic regression for discovering biomarkers of spontaneous preterm birth (SPTB) from gene expression data.
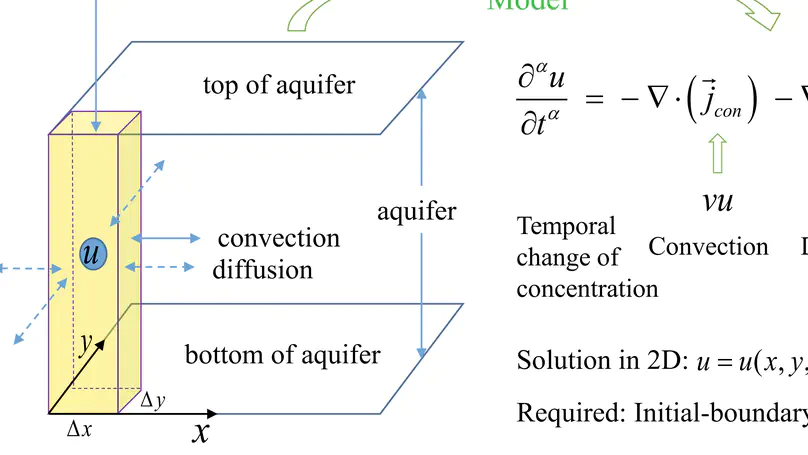
In this work, we provide a compact finite difference scheme (CFDS) of 2D time-fractional convection-diffusion equation (TF-CDE) for solving fluid dynamics problem especially groundwater pollution.
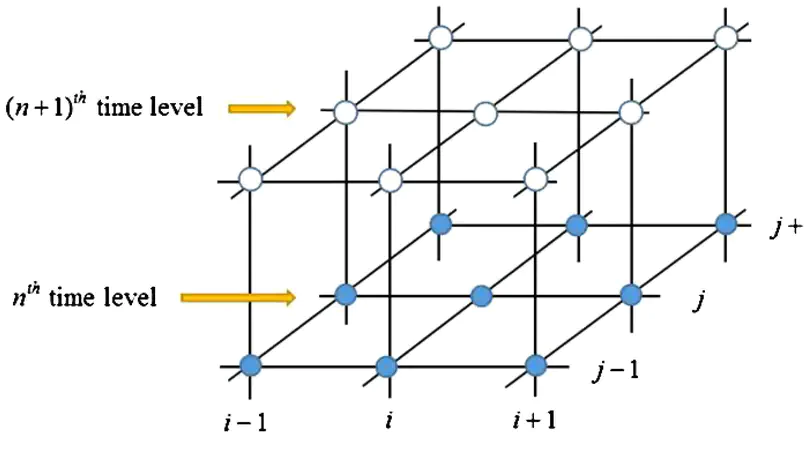
In this work, fourth order compact finite difference scheme of two-dimensional convection diffusion equation to solve groundwater pollution problems
Projects
Contact
- sdnully2012@163.com, lingyuli@hku.hk
- (+86)18366138024, (+852)59186312
- Room 1-05, 5 Sassoon Road, Hong Kong SAR,
- Jockey Club Interdisciplinary Research Building
-
Monday - Friday 09:00 to 19:00
Saturday - Sunday 10:00 to 17:00