Identifying diagnostic biomarkers of breast cancer based on gene expression data and ensemble feature selection
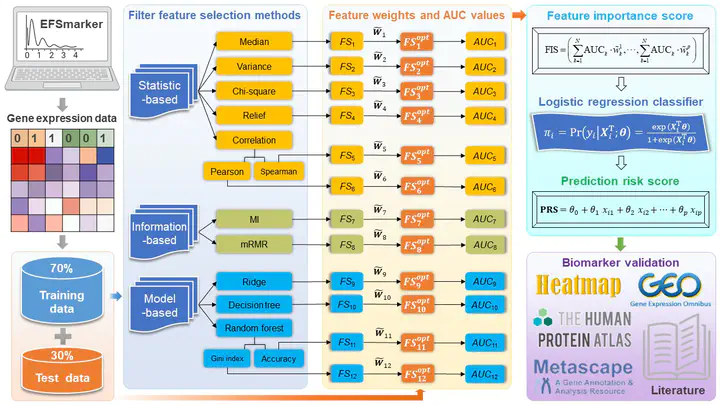 Image credit: Lingyu Li
Image credit: Lingyu Li
Abstract
In recent years, the identification of biomarkers or signatures based on gene expression profiling data has attracted much attention in bioinformatics. The successful discovery of breast cancer (BRCA) biomarkers will be beneficial in reducing the risk of BRCA among patients for early detection. This paper proposes an Ensemble Feature Selection method to screen biomarkers (abbreviated as EFSmarker) for BRCA from publically available gene expression data. Our proposed biomarker discovery strategy not only utilizes the feature contribution but also considers the prediction accuracy simultaneously, which may also serve as a model for identifying unknown biomarkers for other diseases from high-throughput gene expression data. The source code and data are available at https://github.com/zpliulab/EFSmarker.
Supplementary notes can be added here, including code, data, math, and images.