FineST: Contrastive learning integrates histology and spatial transcriptomics for nuclei-resolved ligand-receptor analysis
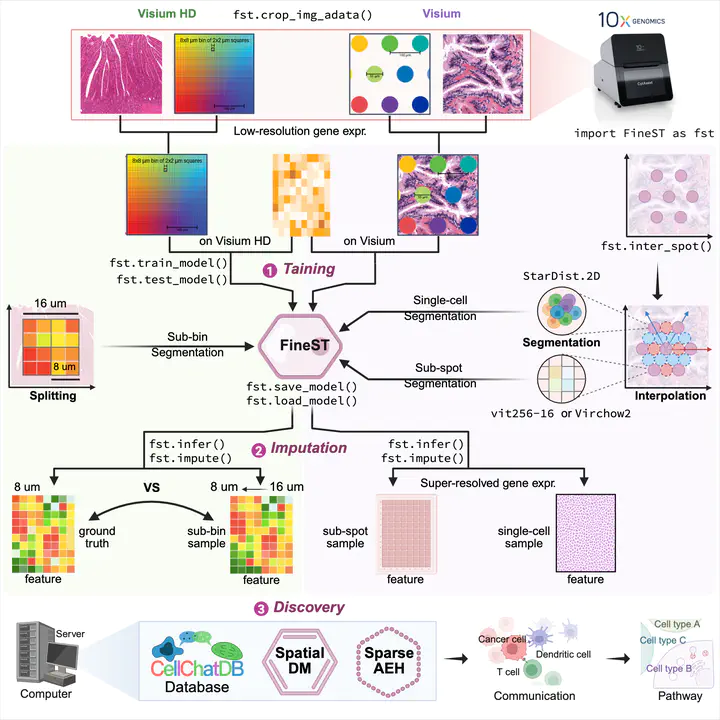 Image credit: [Lingyu Li]
Image credit: [Lingyu Li]
Abstract
Spatial transcriptomics (ST) has emerged as a powerful tool for analyzing cell-cell communication (CCC) across various biological processes, ranging from embryonic development to cancer progression. However, its limited resolution and high data sparsity hinder the detailed characterization of CCC patterns within complex tissues. Here, we introduce FineST, a deep contrastive learning model that leverages a histology foundation model to fuse ST and histology images, enabling Fine-grained Spatial Transcriptomics analysis. This approach facilitates precise nuclei segmentation, high-resolution RNA expression imputation, and the identification of intricate ligand-receptor interactions. Using both colorectal cancer VisiumHD and breast cancer Xenium datasets, we demonstrate that FineST significantly outperforms existing methods in high-resolution RNA imputation, cell type prediction, and CCC pattern discovery. With focused application to the Visium platform, FineST reveals novel biological insights into tumor-immune interactions across multiple cancer types, including invasive fronts in breast cancer, tertiary lymphoid structures in nasopharyngeal carcinoma, and PD-1 therapy resistance barriers in hepatocellular carcinoma. These findings highlight a new paradigm in ST analysis through the integration of readily available histology images.
Supplementary notes can be added here, including code, data, math, and images.