getDNB: identifying dynamic network biomarkers from time-varying gene regulations utilizing graph embedding techniques
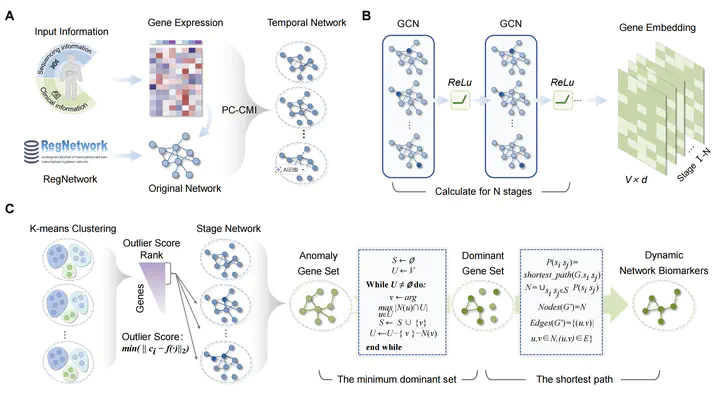 Image credit: [Lingyu Li]
Image credit: [Lingyu Li]
Abstract
Hepatocellular carcinoma (HCC) remains difficult to detect early because conventional diagnostics rely on static biomarkers that emerge late. We present getDNB, a computational framework that identifies dynamic network biomarkers (DNBs) from temporally evolving gene regulatory networks via graph embeddings and anomaly detection. Briefly, getDNB has three steps: (i) construct stage-specific regulatory networks to capture molecular dynamics during HCC progression; (ii) use graph convolutional networks to derive topology-preserving low-dimensional embeddings; and (iii) quantify gene-level abnormalities via K-means clustering and outlier scores, then refine candidates using minimum dominating set and shortest-path criteria to ensure connectivity and reduce redundancy. Moreover, we define the Dynamic Network Index (DNI) to quantify temporal disorder and flag critical transition states. Applied to a real-world HCC dataset, getDNB identified 33 robust DNBs and their interaction network, achieving high predictive accuracy (AUROC = 0.929). Notably, the DNI showed a pronounced increase at the pre-disease stage, consistent with complex systems transition theory. Functional enrichment associated these DNBs with oncogenic pathways, including hepatocellular carcinoma, hepatitis B infection, and cell cycle regulation.
Supplementary notes can be added here, including code, data, math, and images.