NetWalkRank: Cancer driver gene prioritization in multiplex gene regulatory networks by a random walk approach
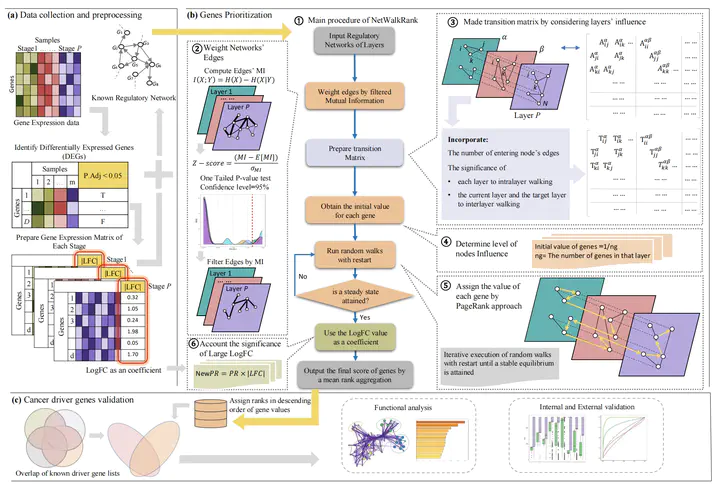 Image credit: [Lingyu Li]
Image credit: [Lingyu Li]
Abstract
Finding and prioritizing cancer driver genes (CDGs) that disrupt normal cell functionality and contribute to cancer occurrence and development is a significant challenge in oncology. Integrating multiple information pertaining to the characteristics of each gene at different stages of the disease and incorporating multiple steps as individual layers in the model provides a more comprehensive understanding of each node or gene. Thus, it is reasonable to organize them into multiplex gene regulatory networks (GRNs). In this work, we present a network-based framework called NetWalkRank, for prioritizing CDGs in the multiplex GRNs with gene expression profiling data. The framework applies the concept of network propagation to calculate the relative impact of each gene in spreading abnormality throughout the multiplex GRNs. It was employed to give priority to the driver genes of hepatocellular carcinoma (HCC) in humans. The performance of NetWalkRank was demonstrated through the ranks and classifications assigned to the known CDGs, which validated its effectiveness. To showcase the predictive capabilities of our proposed framework, we trained a random forest model that utilizes the obtained scores to accurately predict CDGs. We compared the advantage and efficiency of our method with other well-known driver gene ranking methods through numerical experiments. The findings show that the usage of GRNs across various steps of multiplex networks in prioritizing and predicting CDGs is significant, as demonstrated by the efficiency and effectiveness of NetWalkRank. All data and source code used in this study are available at https://github.com/zpliulab/NetWalkRank.
Supplementary notes can be added here, including code, data, math, and images.