Robust biomarker screening from gene expression data by stable machine learning-recursive feature elimination methods
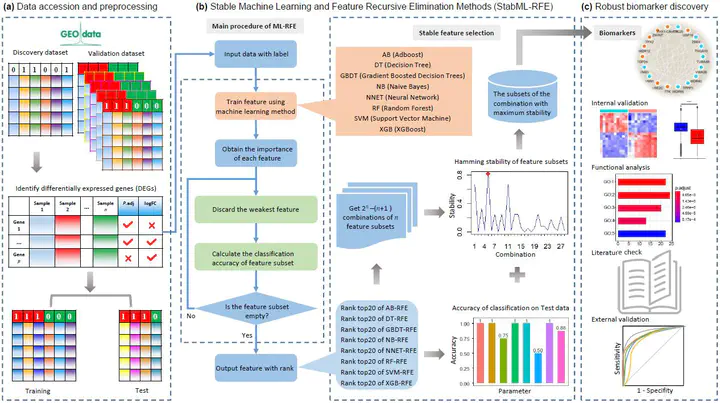 Image credit: Lingyu Li
Image credit: Lingyu Li
摘要
Recently, identifying robust biomarkers or signatures from gene expression profiling data has attracted much attention in computational biomedicine. The successful discovery of biomarkers for complex diseases such as spontaneous preterm birth (SPTB) and high-grade serous ovarian cancer (HGSOC) will be beneficial to reduce the risk of preterm birth and ovarian cancer among women for early detection and intervention. In this paper, we propose a stable machine learning-recursive feature elimination (StabML-RFE for short) strategy for screening robust biomarkers from high-throughput gene expression data. We employ eight popular machine learning methods, namely AdaBoost (AB), Decision Tree (DT), Gradient Boosted Decision Trees (GBDT), Naive Bayes (NB), Neural Network (NNET), Random Forest (RF), Support Vector Machine (SVM) and XGBoost (XGB), to train on all feature genes of training data, apply recursive feature elimination (RFE) to remove the least important features sequentially, and obtain eight gene subsets with feature importance ranking. Then we select the top-ranking features in each ranked subset as the optimal feature subset. We establish a stability metric aggregated with classification performance on test data to assess the robustness of the eight different feature selection techniques. Finally, StabML-RFE chooses the high-frequent features in the subsets of the combination with maximum stability value as robust biomarkers. Particularly, we verify the screened biomarkers not only via internal validation, functional enrichment analysis and literature check, but also via external validation on two real-world SPTB and HGSOC datasets respectively. Obviously, the proposed StabML-RFE biomarker discovery pipeline easily serves as a model for identifying diagnostic biomarkers for other complex diseases from omics data.# Summary. An optional shortened abstract.
Supplementary notes can be added here, including code, data, method, and images.